Southeast Regional Node Meeting – November 17, 2023
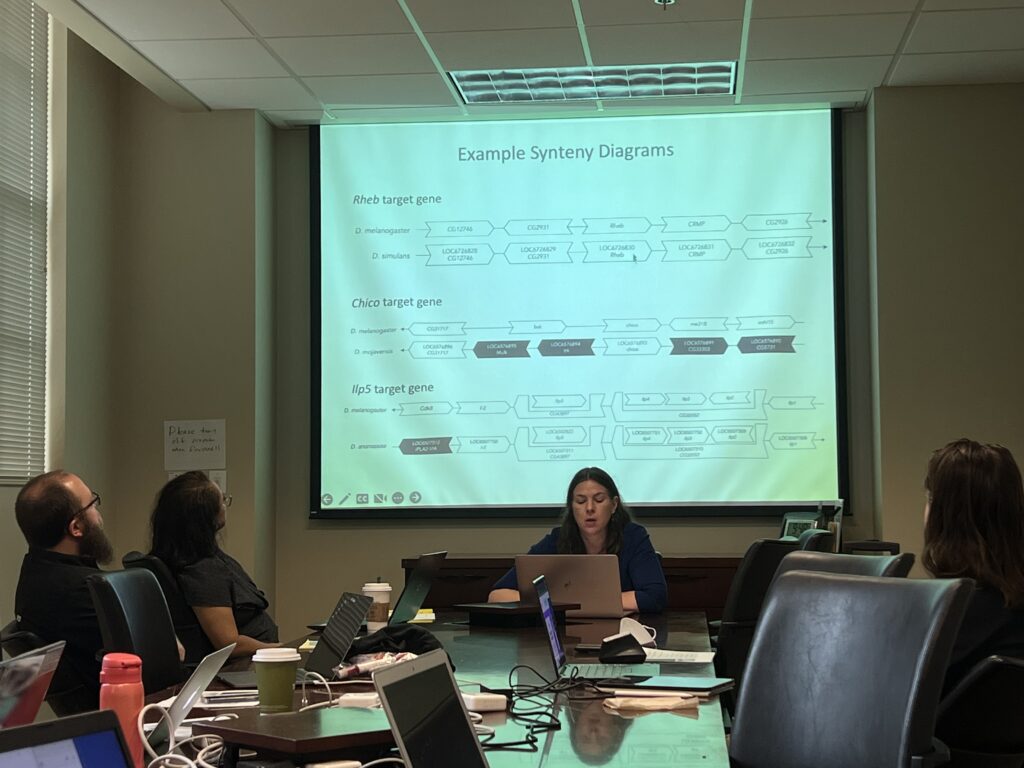
On November 17, 2023, the Southeast Regional Node faculty and students came together for the Fall 2023 Virtual Symposium to highlight excellent student research in gene annotations of F-element and Pathways Projects. In addition, we were very fortunate to have as a keynote speaker Jason Williams of the Cold Spring Harbor Laboratory DNA Learning Center, where he gave some valuable insight into how his career came about in terms of computational biology, bioinformatics, and skills-based instruction and pedagogy with high school and college students. Jason Williams is also the founder of Life Science Trainers, as per a Hudson Alpha article.
Many thanks to everyone for spending their Friday with us to share their work and ideas, we are looking forward to future virtual symposium events for more excellence.
What worked well for your event that might help others plan similar events?
Zoom was used to facilitate the event. Two breakout rooms were set up and students were assigned to one or the other. The schedule was sent out a day or two ahead of time, so that the students knew which breakout room they would be presenting in. Each breakout room was locally recorded and is available to watch on Box (contact Sarah Crocker-Buta for access).
What would your Node do differently based on your experiences?
It remains difficult to limit student presentations and Q&A to 5 minutes because of unexpected computer issues. In the future, Node organizers might need to consider having a moderator receive all presentations and share it on a working laptop for each speaker, in order to mitigate the technical difficulties. One breakout room had finished their presentations approximately 15 minutes before the other, which resulted in the guest speaker waiting in the main room for a period of time before the breakout rooms were closed. It was definitely a learning experience.