NY/NJ Regional Node Meeting – April 25, 2023
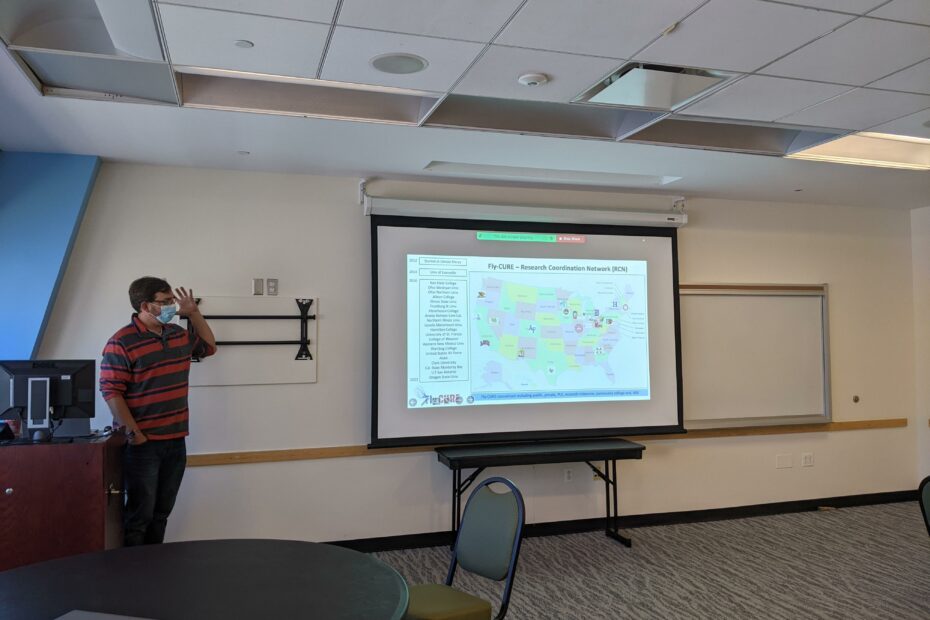
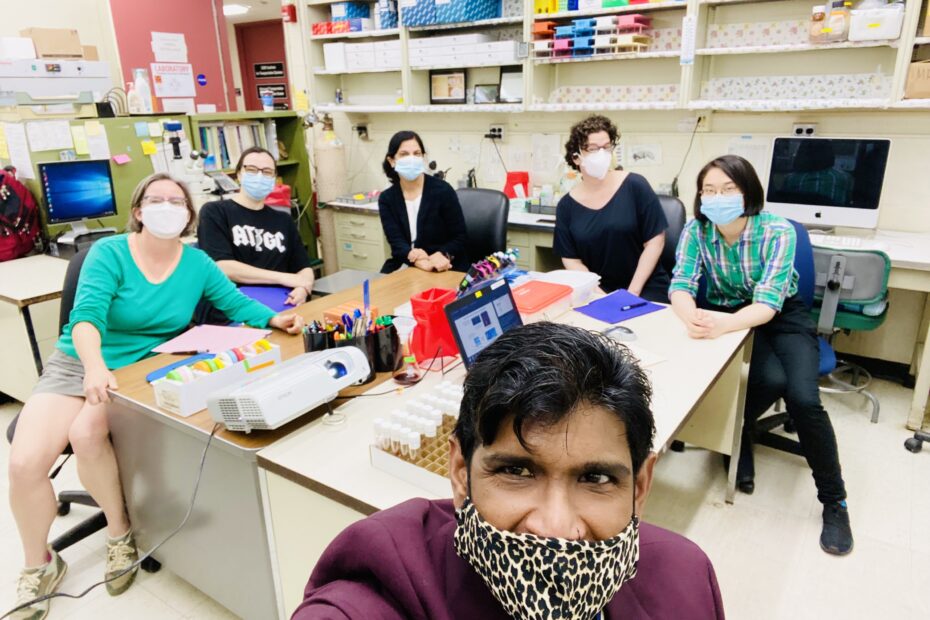
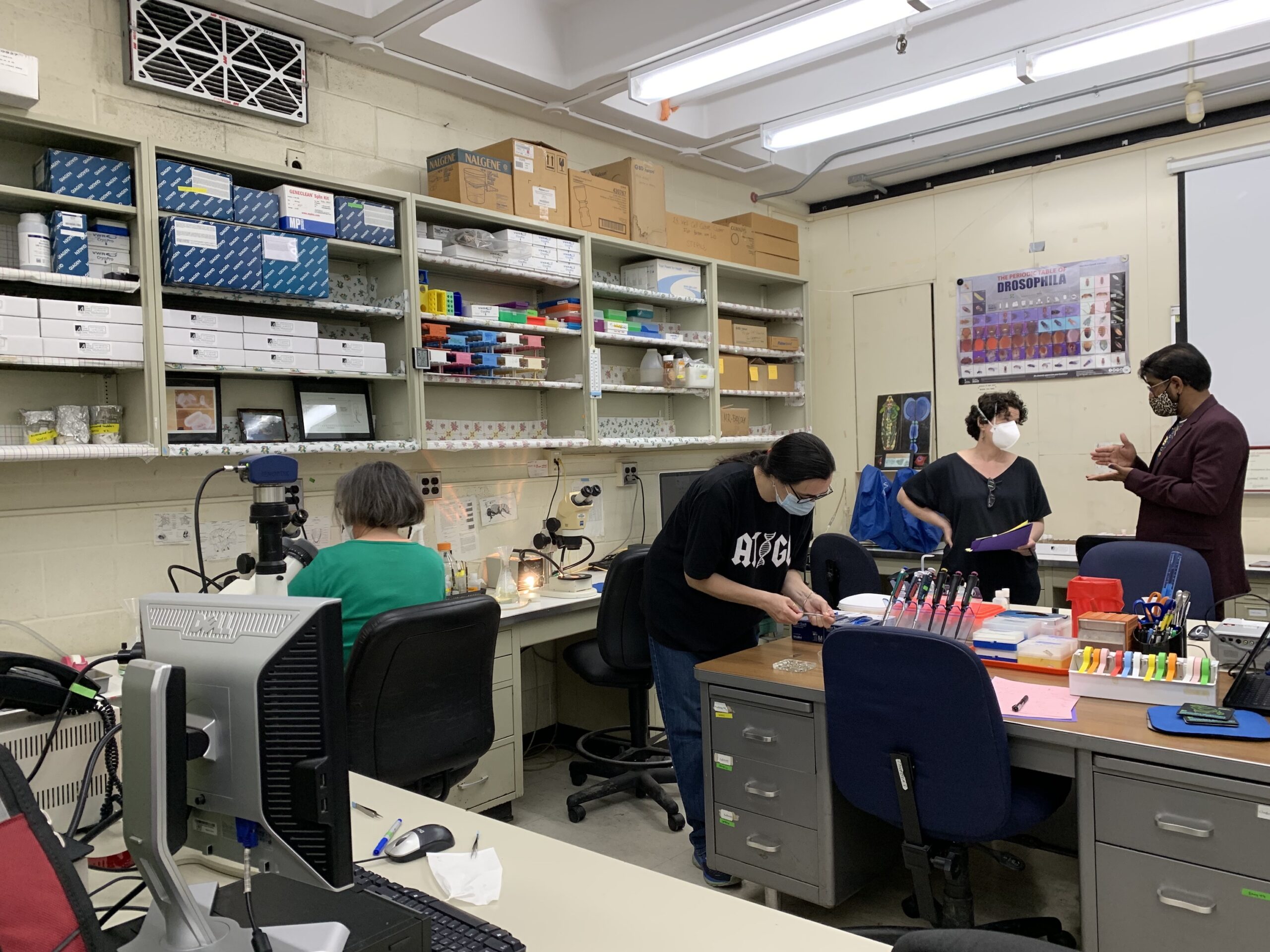
The New York & New Jersey Regional Node held its Spring 2023 professional development meeting virtually with six GEP members and three undergraduate students in attendance. The meeting began with brief introductions followed by three undergraduate student presentations. Alexis Lemberikman (Baruch College) gave a presentation on her work on the Pathways Project (under the mentorship of Dr. Krista Dobi) and two students from Hofstra University (Angad Dev and Margaret Peck) gave presentations on their projects on the Parasitoid Wasps Project (under the mentorship of Dr. Brittany Miller).
Afterwards, students went to a breakout room to meet with each other while the faculty presented short implementation talks. Dr. Brittany Miller gave a short presentation on her experiences implementing the Parasitoid Wasps Project in her bioinformatics class and Dr. Rebecca Spokony talked about how she has used the Understanding Eukaryotic Genes Modules in her developmental biology class. The meeting ended with a discussion of possible Regional Node Events for next academic year. This one and a half hour meeting was an excellent opportunity for members to learn from each other and support the accomplishments of our amazing students.
What worked well for your event that might help others plan similar events?
A survey was sent to the Node well in advance of the event to find out which general dates and times worked for them as well as ask if there were any faculty who wanted to give a short talk or had students who might be interested. This enabled the event organizer to focus on a date and time that worked best for the speakers and members of the Node.
What would your Node do differently based on your experiences?
Organizers discussed offering talks that were not about projects that have already been implemented, but instead about projects that the faculty plan to implement so the rest of the group could give feedback prior to the implementation. This is the plan for next time!