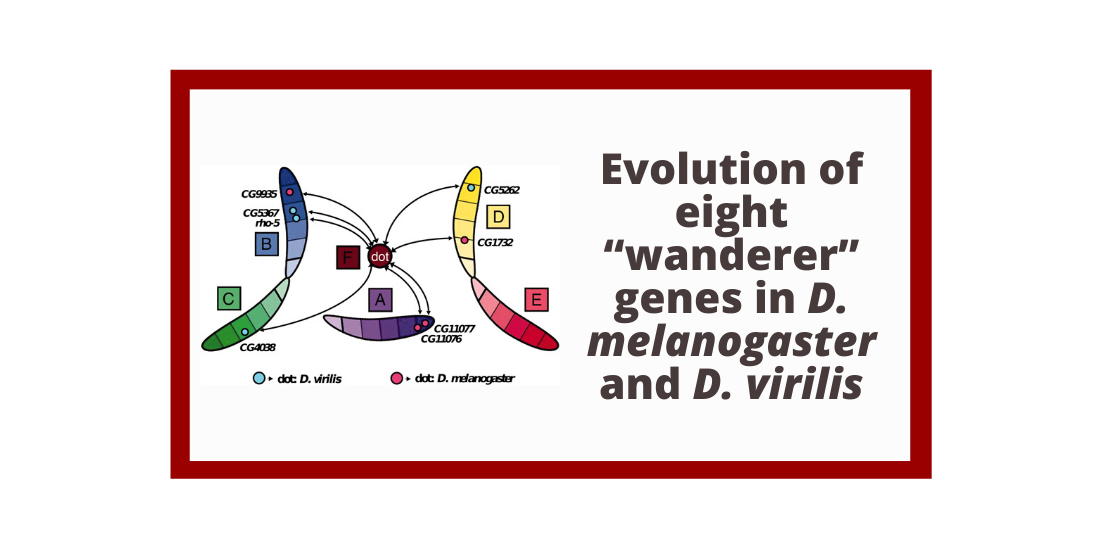
Abstract: The distal arm of the fourth (“dot”) chromosome of Drosophila melanogaster is unusual in that it exhibits an amalgamation of heterochromatic properties (e.g., dense packaging, late replication) and euchromatic properties (e.g., gene density similar to euchromatic domains, replication during polytenization). To examine the evolution of this unusual domain, we undertook a comparative study by generating high-quality sequence data and manually curating gene models for the dot chromosome of D. virilis (Tucson strain 15010-1051.88). Our analysis shows that the dot chromosomes of D. melanogaster and D. virilis have higher repeat density, larger gene size, lower codon bias, and a higher rate of gene rearrangement compared to a reference euchromatic domain. Analysis of eight “wanderer” genes (present in a euchromatic chromosome arm in one species and on the dot chromosome in the other) shows that their characteristics are similar to other genes in the same domain, which suggests that these characteristics are features of the domain and are not required for these genes to function. Comparison of this strain of D. virilis with the strain sequenced by the Drosophila 12 Genomes Consortium (Tucson strain 15010-1051.87) indicates that most genes on the dot are under weak purifying selection. Collectively, despite the heterochromatin-like properties of this domain, genes on the dot evolve to maintain function while being responsive to changes in their local environment.
Leung W, Shaffer CD, Cordonnier T, et al. Evolution of a distinct genomic domain in Drosophila: comparative analysis of the dot chromosome in Drosophila melanogaster and Drosophila virilis. Genetics. 2010;185(4):1519‐1534. doi:10.1534/genetics.110.116129

